Ubiquitin ligase
| Ubiquitin—protein ligase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| EC no. | 2.3.2.27 | ||||||||
| CAS no. | 74812-49-0 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Ubiquitin ligase | |
|---|---|
| Identifiers | |
| Symbol | Ubiquitin ligase |
| OPM superfamily | 471 |
| OPM protein | 4v6p |
| Membranome | 240 |
A ubiquitin ligase (also called an E3 ubiquitin ligase) is a protein that recruits an E2 ubiquitin-conjugating enzyme that has been loaded with ubiquitin, recognizes a protein substrate, and assists or directly catalyzes the transfer of ubiquitin from the E2 to the protein substrate. In simple and more general terms, the ligase enables movement of ubiquitin from a ubiquitin carrier to another protein (the substrate) by some mechanism. The ubiquitin, once it reaches its destination, ends up being attached by an isopeptide bond to a lysine residue, which is part of the target protein.[2] E3 ligases interact with both the target protein and the E2 enzyme, and so impart substrate specificity to the E2. Commonly, E3s polyubiquitinate their substrate with Lys48-linked chains of ubiquitin, targeting the substrate for destruction by the proteasome. However, many other types of linkages are possible and alter a protein's activity, interactions, or localization. Ubiquitination by E3 ligases regulates diverse areas such as cell trafficking, DNA repair, and signaling and is of profound importance in cell biology. E3 ligases are also key players in cell cycle control, mediating the degradation of cyclins, as well as cyclin dependent kinase inhibitor proteins.[3] The human genome encodes over 600 putative E3 ligases, allowing for tremendous diversity in substrates.[4]
Ubiquitination system
[edit]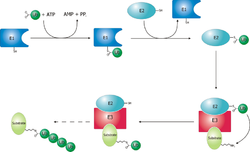
The ubiquitin ligase is referred to as an E3, and operates in conjunction with an E1 ubiquitin-activating enzyme and an E2 ubiquitin-conjugating enzyme. There is one major E1 enzyme, shared by all ubiquitin ligases, that uses ATP to activate ubiquitin for conjugation and transfers it to an E2 enzyme. The E2 enzyme interacts with a specific E3 partner and transfers the ubiquitin to the target protein. The E3, which may be a multi-protein complex, is, in general, responsible for targeting ubiquitination to specific substrate proteins.[citation needed]
The ubiquitylation reaction proceeds in three or four steps depending on the mechanism of action of the E3 ubiquitin ligase. In the conserved first step, an E1 cysteine residue attacks the ATP-activated C-terminal glycine on ubiquitin, resulting in a thioester Ub-S-E1 complex. The energy from ATP and diphosphate hydrolysis drives the formation of this reactive thioester, and subsequent steps are thermoneutral. Next, a transthiolation reaction occurs, in which an E2 cysteine residue attacks and replaces the E1. HECT domain type E3 ligases will have one more transthiolation reaction to transfer the ubiquitin molecule onto the E3, whereas the much more common RING finger domain type ligases transfer ubiquitin directly from E2 to the substrate.[5] The final step in the first ubiquitylation event is an attack from the target protein lysine amine group, which will remove the cysteine, and form a stable isopeptide bond.[6] One notable exception to this is p21 protein, which appears to be ubiquitylated using its N-terminal amine, thus forming a peptide bond with ubiquitin.[7]
Ubiquitin ligase families
[edit]Humans have an estimated 500-1000 E3 ligases, which impart substrate specificity onto the E1 and E2.[8] The E3 ligases are classified into four families: HECT, RING-finger, U-box, and PHD-finger.[8] The RING-finger E3 ligases are the largest family and contain ligases such as the anaphase-promoting complex (APC) and the SCF complex (Skp1-Cullin-F-box protein complex). SCF complexes consist of four proteins: Rbx1, Cul1, Skp1, which are invariant among SCF complexes, and an F-box protein, which varies. Around 70 human F-box proteins have been identified.[9] F-box proteins contain an F-box, which binds the rest of the SCF complex, and a substrate binding domain, which gives the E3 its substrate specificity.[8]
Mono- and poly-ubiquitylation
[edit]
Ubiquitin signaling relies on the diversity of ubiquitin tags for the specificity of its message. A protein can be tagged with a single ubiquitin molecule (monoubiquitylation), or variety of different chains of ubiquitin molecules (polyubiquitylation).[11] E3 ubiquitin ligases catalyze polyubiquitination events much in the same way as the single ubiquitylation mechanism, using instead a lysine residue from a ubiquitin molecule currently attached to substrate protein to attack the C-terminus of a new ubiquitin molecule.[6][11] For example, a common 4-ubiquitin tag, linked through the lysine at position 48 (K48) recruits the tagged protein to the proteasome, and subsequent degradation.[11] However, all seven of the ubiquitin lysine residues (K6, K11, K27, K29, K33, K48, and K63), as well as the N-terminal methionine are used in chains in vivo.[11]
Monoubiquitination has been linked to membrane protein endocytosis pathways. For example, phosphorylation of the Tyrosine at position 1045 in the Epidermal Growth Factor Receptor (EGFR) can recruit the RING type E3 ligase c-Cbl, via an SH2 domain. C-Cbl monoubiquitylates EGFR, signaling for its internalization and trafficking to the lysosome.[12]
Monoubiquitination also can regulate cytosolic protein localization. For example, the E3 ligase MDM2 ubiquitylates p53 either for degradation (K48 polyubiquitin chain), or for nuclear export (monoubiquitylation). These events occur in a concentration dependent fashion, suggesting that modulating E3 ligase concentration is a cellular regulatory strategy for controlling protein homeostasis and localization.[13]
Substrate recognition
[edit]Ubiquitin ligases are the final, and potentially the most important determinant of substrate specificity in ubiquitination of proteins.[14] The ligases must simultaneously distinguish their protein substrate from thousands of other proteins in the cell, and from other (ubiquitination-inactive) forms of the same protein. This can be achieved by different mechanisms, most of which involve recognition of degrons: specific short amino acid sequences or chemical motifs on the substrate.[15]
N-degrons
[edit]Proteolytic cleavage can lead to exposure of residues at the N-terminus of a protein. According to the N-end rule, different N-terminal amino acids (or N-degrons) are recognized to a different extent by their appropriate ubiquitin ligase (N-recognin), influencing the half-life of the protein.[16] For instance, positively charged (Arg, Lys, His) and bulky hydrophobic amino acids (Phe, Trp, Tyr, Leu, Ile) are recognized preferentially and thus considered destabilizing degrons since they allow faster degradation of their proteins.[17]
Phosphodegrons
[edit]
A degron can be converted into its active form by a post-translational modification[19] such as phosphorylation of a tyrosine, serine or threonine residue.[20] In this case, the ubiquitin ligase exclusively recognizes the phosphorylated version of the substrate due to stabilization within the binding site. For example, FBW7, the F-box substrate recognition unit of an SCFFBW7ubiquitin ligase, stabilizes a phosphorylated substrate by hydrogen binding its arginine residues to the phosphate, as shown in the figure to the right. In absence of the phosphate, residues of FBW7 repel the substrate.[18]
Oxygen and small molecule dependent degrons
[edit]The presence of oxygen or other small molecules can influence degron recognition.[18] The von Hippel-Lindau (VHL) protein (substrate recognition part of a specific E3 ligase), for instance, recognizes the hypoxia-inducible factor alpha (HIF-α) only under normal oxygen conditions, when its proline is hydroxylated. Under hypoxia, on the other hand, HIF-a is not hydroxylated, evades ubiquitination and thus operates in the cell at higher concentrations which can initiate transcriptional response to hypoxia.[21] Another example of small molecule control of protein degradation is phytohormone auxin in plants.[22] Auxin binds to TIR1 (the substrate recognition domain of SCFTIR1ubiquitin ligase) increasing the affinity of TIR1 for its substrates (transcriptional repressors: Aux/IAA), and promoting their degradation.
Misfolded and sugar degrons
[edit]In addition to recognizing amino acids, ubiquitin ligases can also detect unusual features on substrates that serve as signals for their destruction.[14] For example, San1 (Sir antagonist 1), a nuclear protein quality control in yeast, has a disordered substrate binding domain, which allows it to bind to hydrophobic domains of misfolded proteins.[14] Misfolded or excess unassembled glycoproteins of the ERAD pathway, on the other hand, are recognized by Fbs1 and Fbs2, mammalian F-box proteins of E3 ligases SCFFbs1and SCFFbs2.[23] These recognition domains have small hydrophobic pockets allowing them to bind high-mannose containing glycans.
Structural motifs
[edit]In addition to linear degrons, the E3 ligase can in some cases also recognize structural motifs on the substrate.[14] In this case, the 3D motif can allow the substrate to directly relate its biochemical function to ubiquitination. This relation can be demonstrated with TRF1 protein (regulator of human telomere length), which is recognized by its corresponding E3 ligase (FBXO4) via an intermolecular beta sheet interaction. TRF1 cannot be ubiquinated while telomere bound, likely because the same TRF1 domain that binds to its E3 ligase also binds to telomeres.[14]
Disease relevance
[edit]E3 ubiquitin ligases regulate homeostasis, cell cycle, and DNA repair pathways, and as a result, a number of these proteins are involved in a variety of cancers, including famously MDM2, BRCA1, and Von Hippel-Lindau tumor suppressor.[24] For example, a mutation of MDM2 has been found in stomach cancer,[25] renal cell carcinoma,[26] and liver cancer[27] (amongst others) to deregulate MDM2 concentrations by increasing its promoter’s affinity for the Sp1 transcription factor, causing increased transcription of MDM2 mRNA.[25] Several proteomics-based experimental techniques are available for identifying E3 ubiquitin ligase-substrate pairs,[28] such as proximity-dependent biotin identification (BioID), ubiquitin ligase-substrate trapping, and tandem ubiquitin-binding entities (TUBEs).
Examples
[edit]- A RING (Really Interesting New Gene) domain binds the E2 conjugase and might be found to mediate enzymatic activity in the E2-E3 complex[29]
- An F-box domain (as in the SCF complex) binds the ubiquitinated substrate. (e.g., Cdc 4, which binds the target protein Sic1; Grr1, which binds Cln).[30]
- A HECT domain, which is involved in the transfer of ubiquitin from the E2 to the substrate.
Individual E3 ubiquitin ligases
[edit]- E3A
- mdm2
- Anaphase-promoting complex (APC)
- UBR5 (EDD1)
- SOCS/ BC-box/ eloBC/ CUL5/ RING
- LNXp80
- CBX4, CBLL1
- HACE1
- HECTD1, HECTD2, HECTD3, HECTD4
- HECW1, HECW2
- HERC1, HERC2, HERC3, HERC4, HERC5, HERC6
- HUWE1, ITCH
- NEDD4, NEDD4L
- PPIL2
- PRPF19
- PIAS1, PIAS2, PIAS3, PIAS4
- RANBP2
- RNF4
- RBX1
- SMURF1, SMURF2
- STUB1
- TOPORS
- TRIP12
- UBE3A, UBE3B, UBE3C, UBE3D
- UBE4A, UBE4B
- UBOX5
- UBR5
- VHL
- WWP1, WWP2
- Parkin
- MKRN1
See also
[edit]References
[edit]- ^ Dou H, Buetow L, Hock A, Sibbet GJ, Vousden KH, Huang DT (January 2012). "Structural basis for autoinhibition and phosphorylation-dependent activation of c-Cbl". Nature Structural & Molecular Biology. 19 (2): 184–92. doi:10.1038/nsmb.2231. PMC 3880865. PMID 22266821.
- ^ Hershko A, Ciechanover A (1998). "The ubiquitin system". Annual Review of Biochemistry. 67: 425–79. doi:10.1146/annurev.biochem.67.1.425. PMID 9759494.
- ^ Teixeira LK, Reed SI (2013). "Ubiquitin ligases and cell cycle control". Annual Review of Biochemistry. 82: 387–414. doi:10.1146/annurev-biochem-060410-105307. PMID 23495935.
- ^ Li W, Bengtson MH, Ulbrich A, Matsuda A, Reddy VA, Orth A, Chanda SK, Batalov S, Joazeiro CA (January 2008). "Genome-wide and functional annotation of human E3 ubiquitin ligases identifies MULAN, a mitochondrial E3 that regulates the organelle's dynamics and signaling". PLOS ONE. 3 (1): e1487. Bibcode:2008PLoSO...3.1487L. doi:10.1371/journal.pone.0001487. PMC 2198940. PMID 18213395.
- ^ Metzger MB, Hristova VA, Weissman AM (February 2012). "HECT and RING finger families of E3 ubiquitin ligases at a glance". Journal of Cell Science. 125 (Pt 3): 531–7. doi:10.1242/jcs.091777. PMC 3381717. PMID 22389392.
- ^ a b Walsh, Christopher (2006). Posttranslational Modification of Proteins: Expanding Nature's Inventory. Englewood, CO: Roberts. ISBN 978-0-9747077-3-0.[page needed]
- ^ Bloom J, Amador V, Bartolini F, DeMartino G, Pagano M (October 2003). "Proteasome-mediated degradation of p21 via N-terminal ubiquitinylation". Cell. 115 (1): 71–82. doi:10.1016/S0092-8674(03)00755-4. PMID 14532004.
- ^ a b c Nakayama KI, Nakayama K (May 2006). "Ubiquitin ligases: cell-cycle control and cancer". Nature Reviews. Cancer. 6 (5): 369–81. doi:10.1038/nrc1881. PMID 16633365. S2CID 19594293.
- ^ Jin J, Cardozo T, Lovering RC, Elledge SJ, Pagano M, Harper JW (November 2004). "Systematic analysis and nomenclature of mammalian F-box proteins". Genes & Development. 18 (21): 2573–80. doi:10.1101/gad.1255304. PMC 525538. PMID 15520277.
- ^ Vijay-Kumar S, Bugg CE, Cook WJ (April 1987). "Structure of ubiquitin refined at 1.8 A resolution". Journal of Molecular Biology. 194 (3): 531–44. doi:10.1016/0022-2836(87)90679-6. PMID 3041007.
- ^ a b c d Behrends C, Harper JW (May 2011). "Constructing and decoding unconventional ubiquitin chains". Nature Structural & Molecular Biology. 18 (5): 520–8. doi:10.1038/nsmb.2066. PMID 21540891. S2CID 19237120.
- ^ Bonifacino JS, Traub LM (2003). "Signals for sorting of transmembrane proteins to endosomes and lysosomes". Annual Review of Biochemistry. 72: 395–447. doi:10.1146/annurev.biochem.72.121801.161800. PMID 12651740.
- ^ Li M, Brooks CL, Wu-Baer F, Chen D, Baer R, Gu W (December 2003). "Mono- versus polyubiquitination: differential control of p53 fate by Mdm2". Science. 302 (5652): 1972–5. Bibcode:2003Sci...302.1972L. doi:10.1126/science.1091362. PMID 14671306. S2CID 43124248.
- ^ a b c d e Zheng N, Shabek N (June 2017). "Ubiquitin Ligases: Structure, Function, and Regulation". Annual Review of Biochemistry. 86 (1): 129–157. doi:10.1146/annurev-biochem-060815-014922. PMID 28375744.
- ^ Ravid T, Hochstrasser M (September 2008). "Diversity of degradation signals in the ubiquitin-proteasome system". Nature Reviews Molecular Cell Biology. 9 (9): 679–90. doi:10.1038/nrm2468. PMC 2606094. PMID 18698327.
- ^ Sriram SM, Kim BY, Kwon YT (October 2011). "The N-end rule pathway: emerging functions and molecular principles of substrate recognition". Nature Reviews Molecular Cell Biology. 12 (11): 735–47. doi:10.1038/nrm3217. PMID 22016057. S2CID 10555455.
- ^ Tasaki T, Sriram SM, Park KS, Kwon YT (2012). "The N-end rule pathway". Annual Review of Biochemistry. 81: 261–89. doi:10.1146/annurev-biochem-051710-093308. PMC 3610525. PMID 22524314.
- ^ a b c Lucas X, Ciulli A (June 2017). "Recognition of substrate degrons by E3 ubiquitin ligases and modulation by small-molecule mimicry strategies" (PDF). Current Opinion in Structural Biology. 44: 101–110. doi:10.1016/j.sbi.2016.12.015. PMID 28130986.
- ^ Herhaus L, Dikic I (September 2015). "Expanding the ubiquitin code through post-translational modification". EMBO Reports. 16 (9): 1071–83. doi:10.15252/embr.201540891. PMC 4576978. PMID 26268526.
- ^ Reinhardt HC, Yaffe MB (September 2013). "Phospho-Ser/Thr-binding domains: navigating the cell cycle and DNA damage response". Nature Reviews Molecular Cell Biology. 14 (9): 563–80. doi:10.1038/nrm3640. PMID 23969844. S2CID 149598.
- ^ Jaakkola P, Mole DR, Tian YM, Wilson MI, Gielbert J, Gaskell SJ, von Kriegsheim A, Hebestreit HF, Mukherji M, Schofield CJ, Maxwell PH, Pugh CW, Ratcliffe PJ (April 2001). "Targeting of HIF-alpha to the von Hippel-Lindau ubiquitylation complex by O2-regulated prolyl hydroxylation". Science. 292 (5516): 468–72. Bibcode:2001Sci...292..468J. doi:10.1126/science.1059796. PMID 11292861. S2CID 20914281.
- ^ Shabek N, Zheng N (April 2014). "Plant ubiquitin ligases as signaling hubs". Nature Structural & Molecular Biology. 21 (4): 293–6. doi:10.1038/nsmb.2804. PMID 24699076. S2CID 41227590.
- ^ Yoshida Y, Mizushima T, Tanaka K (2019-02-19). "Sugar-Recognizing Ubiquitin Ligases: Action Mechanisms and Physiology". Frontiers in Physiology. 10: 104. doi:10.3389/fphys.2019.00104. PMC 6389600. PMID 30837888.
- ^ Lipkowitz S, Weissman AM (August 2011). "RINGs of good and evil: RING finger ubiquitin ligases at the crossroads of tumour suppression and oncogenesis". Nature Reviews. Cancer. 11 (9): 629–43. doi:10.1038/nrc3120. PMC 3542975. PMID 21863050.
- ^ a b Hou YC, Deng JY (January 2015). "Role of E3 ubiquitin ligases in gastric cancer". World Journal of Gastroenterology. 21 (3): 786–93. doi:10.3748/wjg.v21.i3.786. PMC 4299330. PMID 25624711.
- ^ de Martino M, Taus C, Wessely IS, Lucca I, Hofbauer SL, Haitel A, Shariat SF, Klatte T (February 2015). "The T309G murine double minute 2 gene polymorphism is an independent prognostic factor for patients with renal cell carcinoma". DNA and Cell Biology. 34 (2): 107–12. doi:10.1089/dna.2014.2653. PMID 25415135.
- ^ Tang T, Song X, Yang Z, Huang L, Wang W, Tan H (November 2014). "Association between murine double minute 2 T309G polymorphism and risk of liver cancer". Tumour Biology. 35 (11): 11353–7. doi:10.1007/s13277-014-2432-9. PMID 25119589. S2CID 16385927.
- ^ Rayner SL, Morsch M, Molloy MP, Shi B, Chung R, Lee A (July 2019). "Using proteomics to identify ubiquitin ligase-substrate pairs: how novel methods may unveil therapeutic targets for neurodegenerative diseases". Cellular and Molecular Life Sciences. 76 (13): 2499–2510. doi:10.1007/s00018-019-03082-9. PMC 11105231. PMID 30919022. S2CID 85527795.
- ^ Ardley HC, Robinson PA (2005). "E3 ubiquitin ligases". Essays in Biochemistry. 41: 15–30. doi:10.1042/EB0410015. PMID 16250895.
- ^ Bai C, Sen P, Hofmann K, Ma L, Goebl M, Harper JW, Elledge SJ (July 1996). "SKP1 connects cell cycle regulators to the ubiquitin proteolysis machinery through a novel motif, the F-box". Cell. 86 (2): 263–74. doi:10.1016/S0092-8674(00)80098-7. PMID 8706131.
External links
[edit]- Quips article describing E3 Ligase function Archived 2012-11-30 at the Wayback Machine at PDBe
- Ubiquitin-Protein+Ligases at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- EC 6.3.2.19
