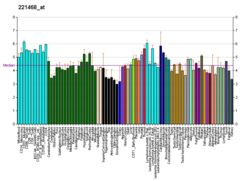
XCR1
The "C" sub-family of chemokine receptors contains only one member: XCR1, the receptor for XCL1 and XCL2 (or lymphotactin-1 and -2).
XCR1 is also known as GPR5.
Function
[edit]The protein encoded by this gene is a chemokine receptor belonging to the G protein-coupled receptor superfamily. The family members are characterized by the presence of 7 transmembrane domains and numerous conserved amino acids. This receptor is most closely related to RBS11 and the MIP1-alpha/RANTES receptor. It transduces a signal by increasing the intracellular calcium ions level. The viral macrophage inflammatory protein-II is an antagonist of this receptor and blocks signaling. Two alternatively spliced transcript variants encoding the same protein have been found for this gene.[5]
Cross-presenting dendritic cells (DCs) in the spleen develop into XCR1+ DCs in the small intestine, T cell zones of Peyer's patches, and T cell zones and sinuses of mesenteric lymph nodes. XCR1+ DCs specialize in cross-presentations of orally applied antigens. The integrin SIRPα is also a differentiating factor for the XCR1+ DCs. The development transcription factor Batf3 helps develop the differences between XCR1+ DCs and CD103+ CD11b- DCs.[6]
XCL1 contributes to chemotaxis only in CD8+ murine cells, but not other DC types, B cells, T cells, or NK cells. Only some of these CD8+ murine cells expressed XCR1 receptors. NK cells release XCL1 along with IFN-γ and some other chemokines upon encountering certain bacteria such as Listeria or MCMV. XCR1+ and CD8+cells work together to cross-present antigen and communicate CD8+ activation. Cross presentation of XCR1+ CD8+ and XCR1+ CD8- cells was strongest, as is expected since they have XCR1 receptors. CD4+ and CD8+ may become outdated terms, since the activity of the cell appears to be primarily dependent upon the expression of XCR1, which will make a population far more similar than the expression of CD4 or CD8.[7]
XCR1+ cells are dependent on the growth factor Ftl3 ligand and are nonexistent in Batf3- deficient mice. Also, XCR1+ DCs are related to CD103+CD11b- DCs.[6]
XCL1 is expressed by medullary thymic epithelial T cells (mTECs) while XCR1 is expressed by thymic dendritic cells (tDCs). This communication helps with the destruction of cells that are not self-tolerant. When mice lose the ability to express XCL1, they are deficient in accumulation of tDCs and producing naturally occurring regulatory T cells (nT reg cells). The displaying of XCL1 by mTECs, tDC chemotaxis, and nT reg cell production are all decreased in mice that lack Aire, demonstrating it as an important regulator of XCL1 production.[8]
Naive CD8+ T cells are prepared when tumors form by cross-presentation via XCR1+ DCs and as a result will require a lower threshold to respond to antigen. Memory CD8+ T lymphocytes (mCTLs) are activated first after infection and then are signaled by CXCR3, IL-12, and CXCL9 by other XCR1+ DCs. In order to make a powerful secondary infection response, cytokine and chemokine signaling between XCR1+ DCs and NK cells must occur.[9]
References
[edit]- ^ a b c GRCh38: Ensembl release 89: ENSG00000173578 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000060509 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Entrez Gene: XCR1 chemokine (C motif) receptor 1".
- ^ a b Becker M, Güttler S, Bachem A, Hartung E, Mora A, Jäkel A, Hutloff A, Henn V, Mages HW, Gurka S, Kroczek RA (2014). "Ontogenic, Phenotypic, and Functional Characterization of XCR1(+) Dendritic Cells Leads to a Consistent Classification of Intestinal Dendritic Cells Based on the Expression of XCR1 and SIRPα". Frontiers in Immunology. 5: 326. doi:10.3389/fimmu.2014.00326. PMC 4112810. PMID 25120540.
- ^ Kroczek RA, Henn V (2012). "The Role of XCR1 and its Ligand XCL1 in Antigen Cross-Presentation by Murine and Human Dendritic Cells". Frontiers in Immunology. 3: 14. doi:10.3389/fimmu.2012.00014. PMC 3342032. PMID 22566900.
- ^ Lei Y, Ripen AM, Ishimaru N, Ohigashi I, Nagasawa T, Jeker LT, Bösl MR, Holländer GA, Hayashi Y, Malefyt Rde W, Nitta T, Takahama Y (February 2011). "Aire-dependent production of XCL1 mediates medullary accumulation of thymic dendritic cells and contributes to regulatory T cell development". The Journal of Experimental Medicine. 208 (2): 383–94. doi:10.1084/jem.20102327. PMC 3039864. PMID 21300913.
- ^ Alexandre YO, Ghilas S, Sanchez C, Le Bon A, Crozat K, Dalod M (January 2016). "XCR1+ dendritic cells promote memory CD8+ T cell recall upon secondary infections with Listeria monocytogenes or certain viruses". The Journal of Experimental Medicine. 213 (1): 75–92. doi:10.1084/jem.20142350. PMC 4710197. PMID 26694969.
External links
[edit]- "Chemokine Receptors: XCR1". IUPHAR Database of Receptors and Ion Channels. International Union of Basic and Clinical Pharmacology.
Further reading
[edit]- Maghazachi AA (June 1999). "Intracellular signalling pathways induced by chemokines in natural killer cells". Cellular Signalling. 11 (6): 385–90. doi:10.1016/S0898-6568(99)00008-X. PMID 10400311.
- Gao JL, Kuhns DB, Tiffany HL, McDermott D, Li X, Francke U, Murphy PM (May 1993). "Structure and functional expression of the human macrophage inflammatory protein 1 alpha/RANTES receptor". The Journal of Experimental Medicine. 177 (5): 1421–7. doi:10.1084/jem.177.5.1421. PMC 2191019. PMID 7683036.
- Heiber M, Docherty JM, Shah G, Nguyen T, Cheng R, Heng HH, Marchese A, Tsui LC, Shi X, George SR (January 1995). "Isolation of three novel human genes encoding G protein-coupled receptors". DNA and Cell Biology. 14 (1): 25–35. doi:10.1089/dna.1995.14.25. PMID 7832990.
- Yoshida T, Imai T, Kakizaki M, Nishimura M, Takagi S, Yoshie O (June 1998). "Identification of single C motif-1/lymphotactin receptor XCR1". The Journal of Biological Chemistry. 273 (26): 16551–4. doi:10.1074/jbc.273.26.16551. PMID 9632725.
- Shan L, Qiao X, Oldham E, Catron D, Kaminski H, Lundell D, Zlotnik A, Gustafson E, Hedrick JA (February 2000). "Identification of viral macrophage inflammatory protein (vMIP)-II as a ligand for GPR5/XCR1". Biochemical and Biophysical Research Communications. 268 (3): 938–41. doi:10.1006/bbrc.2000.2235. PMID 10679309.
- Maho A, Bensimon A, Vassart G, Parmentier M (2000). "Mapping of the CCXCR1, CX3CR1, CCBP2 and CCR9 genes to the CCR cluster within the 3p21.3 region of the human genome". Cytogenetics and Cell Genetics. 87 (3–4): 265–8. doi:10.1159/000015443. PMID 10702689. S2CID 1178132.
- Kurt RA, Bauck M, Harma S, McCulloch K, Baher A, Urba WJ (May 2001). "Role of C chemokine lymphotactin in mediating recruitment of antigen-specific CD62L(lo) cells in vitro and in vivo". Cellular Immunology. 209 (2): 83–8. doi:10.1006/cimm.2001.1790. PMID 11446740.
- Shinkai H, Morozumi T, Toki D, Eguchi-Ogawa T, Muneta Y, Awata T, Uenishi H (April 2005). "Genomic structure of eight porcine chemokine receptors and intergene sharing of an exon between CCR1 and XCR1". Gene. 349: 55–66. doi:10.1016/j.gene.2004.10.017. PMID 15777643.
- Lüttichau HR, Johnsen AH, Jurlander J, Rosenkilde MM, Schwartz TW (June 2007). "Kaposi sarcoma-associated herpes virus targets the lymphotactin receptor with both a broad spectrum antagonist vCCL2 and a highly selective and potent agonist vCCL3". The Journal of Biological Chemistry. 282 (24): 17794–805. doi:10.1074/jbc.M702001200. PMID 17403668.
External links
[edit]- XCR1+receptor,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
This article incorporates text from the United States National Library of Medicine, which is in the public domain.