Ankyrin repeat
This article possibly contains original research. (July 2015) |
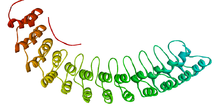
| Ankyrin repeat domain | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||
| Identifiers | |||||||||||
| Symbol | Ank | ||||||||||
| Pfam | PF00023 | ||||||||||
| InterPro | IPR002110 | ||||||||||
| SMART | SM00248 | ||||||||||
| PROSITE | PDOC50088 | ||||||||||
| SCOP2 | 1awc / SCOPe / SUPFAM | ||||||||||
| |||||||||||
The ankyrin repeat is a 33-residue motif in proteins consisting of two alpha helices separated by loops, first discovered in signaling proteins in yeast Cdc10 and Drosophila Notch. Domains consisting of ankyrin tandem repeats mediate protein–protein interactions and are among the most common structural motifs in known proteins. They appear in bacterial, archaeal, and eukaryotic proteins, but are far more common in eukaryotes. Ankyrin repeat proteins, though absent in most viruses, are common among poxviruses. Most proteins that contain the motif have four to six repeats, although its namesake ankyrin contains 24, and the largest known number of repeats is 34, predicted in a protein expressed by Giardia lamblia.[2]
Ankyrin repeats typically fold together to form a single, linear solenoid structure called ankyrin repeat domains. These domains are one of the most common protein–protein interaction platforms in nature. They occur in a large number of functionally diverse proteins, mainly from eukaryotes. The few known examples from prokaryotes and viruses may be the result of horizontal gene transfers.[3] The repeat has been found in proteins of diverse function such as transcriptional initiators, cell cycle regulators, cytoskeletal, ion transporters, and signal transducers. The ankyrin fold appears to be defined by its structure rather than its function, since there is no specific sequence or structure that is universally recognised by it.
Considering the atomic structures of individual ankyrin repeats, the loop is often a type 1 beta bulge loop, while both alpha-helices commonly have a Schellman loop at their N-terminus.
Role in protein folding
[edit]The ankyrin-repeat sequence motif has been studied using multiple sequence alignment to determine conserved amino acid residues critical for folding and stability. The residues on the wide lateral surface of ankyrin repeat structures are variable, often hydrophobic, and involved mainly in mediating protein–protein interactions. An artificial protein design based on a consensus sequence derived from sequence alignment has been synthesized and found to fold stably, representing the first designed protein with multiple repeats.[4] More extensive design strategies have used combinatorial sequences to "evolve" ankyrin-repeats that recognize particular protein targets, a technique that has been presented as an alternative to antibody design for applications requiring high-affinity binding.[5] A structure-based study involving a range of ankyrin proteins of known structures, shows that consensus-based ankyrin proteins are very stable since they maximize the energetic gap between the folding and unfolding structures, encoding a densely connected network of favourable interactions among conserved sequence motifs, like the TPLX motif.[6] The same study shows that insertions in the canonical framework of ankyrin repeats are enriched in conflictive interactions, that are related to function. The same applies to interactions surrounding deletion hotspots. These might be related to complex folding/unfolding transitions that are important to the partner recognition and interaction.
Ankyrin-repeat proteins present an unusual problem in the study of protein folding, which has largely focused on globular proteins that form well-defined tertiary structure stabilized by long-range, nonlocal residue-residue contacts. Ankyrin repeats, by contrast, contain very few such contacts (that is, they have a low contact order). Most studies have found that ankyrin repeats fold in a two-state folding mechanism, suggesting a high degree of folding cooperativity despite the local inter-residue contacts and the evident need for successful folding with varying numbers of repeats. Some evidence, based on synthesis of truncated versions of natural repeat proteins,[7] and on the examination of phi values,[8] suggests that the C-terminus forms the folding nucleation site.
Clinical significance
[edit]Ankyrin-repeat proteins have been associated with a number of human diseases. These proteins include the cell cycle inhibitor p16, which is associated with cancer, and the Notch protein (a key component of cell signalling pathways) which can cause the neurological disorder CADASIL when the repeat domain is disrupted by mutations.[2]
A specialized family of ankyrin proteins known as muscle ankyrin repeat proteins (MARPs) are involved with the repair and regeneration of muscle tissue following damage due to injury and stress.[9]
A natural variation between glutamine and lysine at position 703 in the 11th ankyrin repeat of ANKK1, known as the TaqI A1 allele,[10] has been credited with encouraging addictive behaviours such as obesity, alcoholism, nicotine dependency and the Eros love style[citation needed] while discouraging juvenile delinquency and neuroticism-anxiety.[11] [failed verification] The variation may affect the specificity of protein interactions made by the ANKK1 protein kinase through this repeat[citation needed].
Human proteins containing this repeat
[edit]ABTB1; ABTB2; ACBD6; ACTBL1; ANK1; ANK2; ANK3; ANKAR; ANKDD1A; ANKEF1; ANKFY1; ANKHD1; ANKIB1; ANKK1; ANKMY1; ANKMY2; ANKRA2; ANKRD1; ANKRD10; ANKRD11; ANKRD12; ANKRD13; ANKRD13A; ANKRD13B; ANKRD13C; ANKRD13D; ANKRD15; ANKRD16; ANKRD17; ANKRD18A; ANKRD18B; ANKRD19; ANKRD2; ANKRD20A1; ANKRD20A2; ANKRD20A3; ANKRD20A4; ANKRD21; ANKRD22; ANKRD23; ANKRD24; ANKRD25; ANKRD26; ANKRD27; ANKRD28; ANKRD30A; ANKRD30B; ANKRD30BL; ANKRD32; ANKRD33; ANKRD35; ANKRD36; ANKRD36B; ANKRD37; ANKRD38; ANKRD39; ANKRD40; ANKRD41; ANKRD42; ANKRD43; ANKRD44; ANKRD45; ANKRD46; ANKRD47; ANKRD49; ANKRD50; ANKRD52; ANKRD53; ANKRD54; ANKRD55; ANKRD56; ANKRD57; ANKRD58; ANKRD60; ANKRD6; ANKRD7; ANKRD9; ANKS1A; ANKS3; ANKS4B; ANKS6; ANKZF1; ASB1; ASB10; ASB11; ASB12; ASB13; ASB14; ASB15; ASB16; ASB2; ASB3; ASB4; ASB5; ASB6; ASB7; ASB8; ASB9; ASZ1; BARD1; BAT4; BAT8; BCL3; BCOR; BCORL1; BTBD11; CAMTA1; CAMTA2; CASKIN1; CASKIN2; CCM1; CDKN2A; CDKN2B; CDKN2C; CDKN2D; CENTB1; CENTB2; CENTB5; CENTG1; CENTG2; CENTG3; CLIP3; CLIP4; CLPB; CTGLF1; CTGLF2; CTGLF3; CTGLF4; CTGLF5; CTTNBP2; DAPK1; DDEF1; DDEF2; DDEFL1; DGKI; DGKZ; DP58; DYSFIP1; DZANK; EHMT1; EHMT2; ESPN; FANK1; FEM1A; FEM1B; GABPB2; GIT1; GIT2; GLS; GLS2; HACE1; HECTD1; IBTK; ILK; INVS; KIDINS220; KRIT1; LRRK1; MAIL; MIB1; MIB2; MPHOSPH8; MTPN; MYO16; NFKB1; NFKB2; NFKBIA; NFKBIB; NFKBIE; NFKBIL1; NFKBIL2; NOTCH1; NOTCH2; NOTCH3; NOTCH4; NRARP; NUDT12; OSBPL1A; OSTF1; PLA2G6; POTE14; POTE15; POTE8; PPP1R12A; PPP1R12B; PPP1R12C; PPP1R13B; PPP1R13L; PPP1R16A; PPP1R16B; PSMD10; RAI14; RFXANK; RIPK4; RNASEL; SHANK1; SHANK2; SHANK3; SNCAIP; TA-NFKBH; TEX14; TNKS; TNKS2; TNNI3K; TP53BP2; TRP7; TRPA1; TRPC3; TRPC4; TRPC5; TRPC6; TRPC7; TRPV1; TRPV2; TRPV3; TRPV4; TRPV5; TRPV6; UACA; USH1G; ZDHHC13; ZDHHC17;
See also
[edit]- DARPin (designed ankyrin repeat protein), an engineered antibody mimetic based on the structure of ankyrin repeats
References
[edit]- ^ PDB: 1N11; Michaely P, Tomchick DR, Machius M, Anderson RG (December 2002). "Crystal structure of a 12 ANK repeat stack from human ANK1". EMBO J. 21 (23): 6387–96. doi:10.1093/emboj/cdf651. PMC 136955. PMID 12456646.
- ^ a b Mosavi L, Cammett T, Desrosiers D, Peng Z (2004). "The ankyrin repeat as molecular architecture for protein recognition". Protein Sci. 13 (6): 1435–48. doi:10.1110/ps.03554604. PMC 2279977. PMID 15152081. Archived from the original on 2004-09-07.
- ^ Bork P (December 1993). "Hundreds of ankyrin-like repeats in functionally diverse proteins: mobile modules that cross phyla horizontally?". Proteins. 17 (4): 363–74. doi:10.1002/prot.340170405. PMID 8108379. S2CID 35224626.
- ^ Mosavi LK, Minor DL, Peng ZY (Dec 2002). "Consensus-derived structural determinants of the ankyrin repeat motif". Proc Natl Acad Sci USA. 99 (25): 16029–34. Bibcode:2002PNAS...9916029M. doi:10.1073/pnas.252537899. PMC 138559. PMID 12461176.
- ^ Binz HK, Amstutz P, Kohl A, et al. (May 2004). "High-affinity binders selected from designed ankyrin repeat protein libraries". Nat. Biotechnol. 22 (5): 575–82. doi:10.1038/nbt962. PMID 15097997. S2CID 1191035.
- ^ Parra RG, Espada R, Verstraete N, Ferreiro DU, et al. (Dec 2015). "Structural and Energetic Characterization of the Ankyrin Repeat Protein Family". PLOS Comput. Biol. 12 (11): 575–82. Bibcode:2015PLSCB..11E4659P. doi:10.1371/journal.pcbi.1004659. PMC 4687027. PMID 26691182.
- ^ Zhang B, Peng Z (Jun 2000). "A minimum folding unit in the ankyrin repeat protein p16(INK4)". J Mol Biol. 299 (4): 1121–32. doi:10.1006/jmbi.2000.3803. PMID 10843863.
- ^ Tang KS, Fersht AR, Itzhaki LS (Jan 2003). "Sequential unfolding of ankyrin repeats in tumor suppressor p16". Structure. 11 (1): 67–73. doi:10.1016/S0969-2126(02)00929-2. PMID 12517341.
- ^ Miller MK, Bang ML, Witt CC, et al. (Nov 2003). "The muscle ankyrin repeat proteins: CARP, ankrd2/Arpp and DARP as a family of titin filament-based stress response molecules". J Mol Biol. 333 (5): 951–64. doi:10.1016/j.jmb.2003.09.012. PMID 14583192.
- ^ Neville MJ, Johnstone EC, Walton RT (Jun 2004). "Identification and characterization of ANKK1: a novel kinase gene closely linked to DRD2 on chromosome band 11q23.1". Hum. Mutat. 23 (6): 540–5. doi:10.1002/humu.20039. PMID 15146457. S2CID 22242611.
- ^ "NCBI Gene summary for DRD2". (interim reference)
External links
[edit]- Eukaryotic Linear Motif resource motif class LIG_TNKBM_1
- Ankyrin+repeat at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
